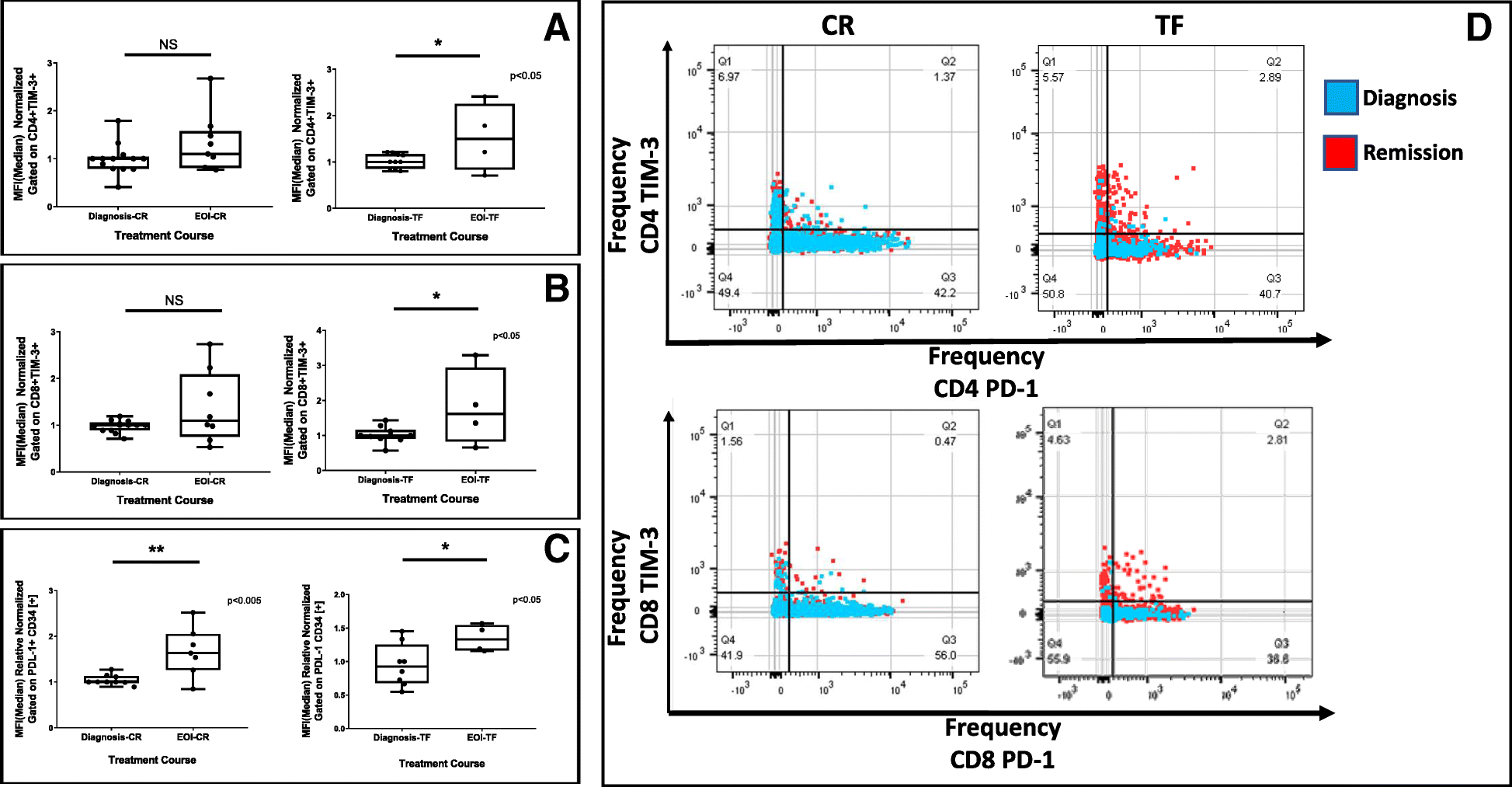
There are two options for constraining the CVs of the G0/G1 and/or G2/M populations. However, if the data does not match the default assumptions, you can constrain the coefficients of variation (CV) for each peak, the peak means, the ratio of the mean peaks, or any combination thereof. If the data fits the models default assumptions of a large identifiableG0/G1 peak, a smaller but identifiable G2/M peak, and little noise, the modeling process will generally create a credible result with no further effort. More details regarding the univariate statistics are available in the univariate statistics link. The ability to model a synchronous S-phase is included as an optional component in the user interface. This modification gives the DJF model the ability to properly fit a synchronous population with a complex S phase distribution. The Fox modification is to make the fit the addition of a Gaussian distribution and the polynomial. The Dean-Jett model fit a second order polynomial (f (x) = Ax^2 + Bx + C) to the S phase. The difference occurs in the fit of the S phase. The G0/G1 and G2/M curves are fit using the same process as the Watson model. The width is estimated in the same manner and a second Gaussian distribution is fit to the data using the same minimization process. Theoretically the G2/M peak will have a mean twice as bright as the G0/G1, but in practice there is some loss in the process and the G2/M peak ends up being not quite twice as bright.
Once the G0/G1 peak is fit, the G2/M mean is placed at 1.75 x the intensity of the G0/G1 mean. The range is unbalanced to the left since little data is expected to occur below the G0/G1 peak, while the S phase cells are expected to occur to the right and are more likely to overlap and skew the fit. A minimization process (least squares fitting) is then executed over a range of -3 to 1 standard deviations about the first guess mean to improve the fit. The standard deviation (SD) or width of the population is then approximated by finding the width of the distribution at 60% of the maximum height. The model initializes by approximating G0/G1 peak as a Gaussian distribution and making an initial guess of the mean by finding the channel with the most cell in the left portion of the data. It assumes only that the data within the G0/G1 and G2/M peaks are normally distributed and that one of those two peaks is identifiable. The Watson model was published by James Watson and colleagues in 1987. It is good laboratory practice to consistently use the same model throughout a study when reporting or publishing statistics. The methods the models employ to calculate their statistics are described below. Consequently, results from one model may vary quite significantly from the other. The two models differ in their mathematical calculations of each phase of the cell cycle. The univariate model will appear by default as shown in Figure 1.įlowJo v10 provides two univariate cell cycle platforms, the Watson Pragmatic algorithm (1) and Dean Jett Fox (DJF) (2). To launch the univariate cell cycle model click on the population of interest in the workspace, then select the Cell Cycle task from the Biology Band. Univariate modeling can be used to create a fit to cell cycle data based on statistics in one dimension, traditionally DNA content.įlowJo provides a simple interface to performing fairly sophisticated DNA/Cell Cycle analysis.
